
Query Expansion for Biomedical Research: A Guide
Struggling to find relevant biomedical research papers? Query expansion can help.
Query expansion improves search results by adding related terms to your query, addressing issues like "vocabulary mismatch" in platforms like PubMed. This is especially useful in biomedical research, where terms like "tumor" and "neoplasms" or "heart attack" and "myocardial infarction" often mean the same thing but are indexed differently.
Here’s a quick overview of the main methods:
- Ontology-Based Expansion: Uses medical databases like UMLS and MeSH to map terms to standardized concepts.
- Neural Embedding Techniques: Leverages machine learning to find deeper semantic relationships between terms.
- LLM-Based Generative Methods: Uses large language models to generate expanded queries, though grounding these in trusted sources is key to avoid inaccuracies.
Each method has its strengths and limitations, but combining them often yields better results. For example, mixing ontology-based and neural approaches has improved retrieval performance by over 20%.
To implement query expansion effectively:
- Preprocess your queries (e.g., clean and tokenize).
- Expand terms using ontology tools like MetaMap or PubMed's MeSH explosion feature.
- Merge sparse (BM25) and dense (BioBERT) retrieval models for optimal results.
Tools like Sourcely and PubMed APIs can further simplify this process, offering features like deep searches and programmatic query expansions. Metrics like MAP and F-measure help evaluate the effectiveness of these techniques.
Whether you're conducting systematic reviews or just trying to find relevant papers, query expansion can make your search process more efficient and accurate.
Build a Medical RAG App using BioMistral, Qdrant, and Llama.cpp
sbb-itb-f7d34da
Main Query Expansion Techniques for Biomedical Research
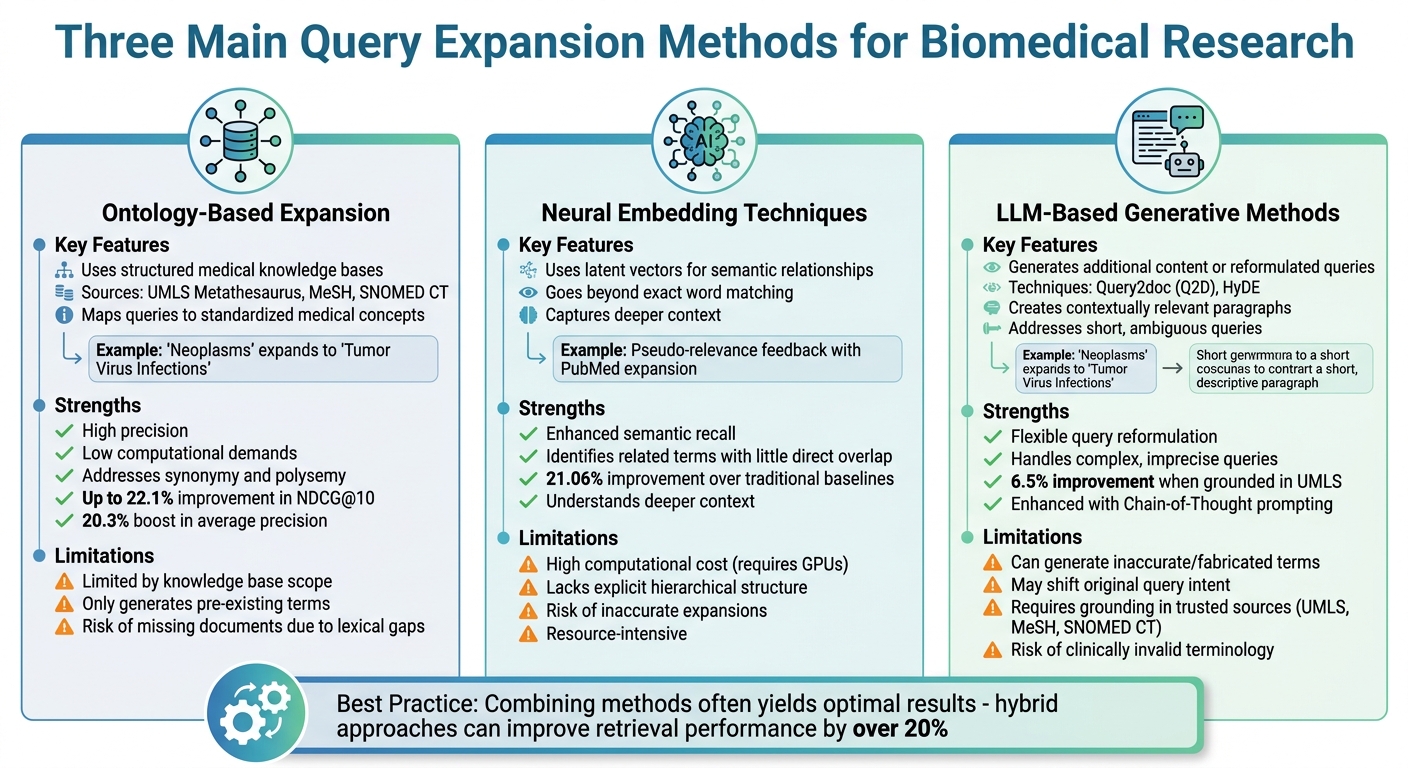
Comparison of Three Main Query Expansion Methods for Biomedical Research
Biomedical researchers use three main strategies to expand search queries: ontology-based methods, neural embedding techniques, and LLM-based generative approaches. These methods aim to address vocabulary mismatches in biomedical searches by carefully balancing precision and recall.
Ontology-Based Expansion
This approach uses structured medical knowledge bases, such as UMLS Metathesaurus, MeSH, and SNOMED CT, to map user queries to standardized medical concepts. It’s particularly effective at addressing synonymy (different terms for the same concept) and polysemy (a single term with multiple meanings).
For instance, the "MeSH explosion" feature automatically expands a broad term like "Neoplasms" to include more specific related terms, such as "Tumor Virus Infections."
Modern tools like BMQExpander take this further by extracting key terms, mapping them to Concept Unique Identifiers (CUIs), and building semantic graphs based on relationships like "has child" or "has parent." Research has shown that this framework can improve retrieval effectiveness (NDCG@10) by up to 22.1% compared to sparse baselines. Additionally, ontology-based term re-weighting has boosted average precision by as much as 20.3% when integrated with search engines like Lucene.
"The fundamental issue [in biomedical literature searching] is synonymy and polysemy", an IEEE Conference Publication explains.
However, ontology-based methods are limited by the scope of their underlying knowledge bases, as they can only generate terms already present in these resources.
Now, let’s look at how neural and embedding-based methods complement these structured techniques.
Neural and Embedding-Based Techniques
These methods rely on latent vectors to capture semantic relationships, moving beyond exact word matching. By understanding deeper context, they can identify related terms even when there is little direct overlap in wording.
For example, combining pseudo-relevance feedback with PubMed-specific expansion has been shown to outperform traditional baselines by 21.06%. Despite their strengths, neural methods require substantial computational resources, such as GPUs, and lack the explicit hierarchical structure provided by ontologies.
Ontology-based methods excel in precision with relatively low computational demands, while neural approaches enhance semantic recall but at a higher computational cost. Ontology-based methods risk missing documents due to lexical gaps, while neural techniques face challenges with inaccurate expansions.
Building on these strengths, LLM-based generative methods offer a different approach to query reformulation. These advanced natural language processing techniques are transforming how researchers interact with literature.
LLM-Based Generative Methods
LLM-based methods enhance queries by generating additional content or reformulated queries with semantically related terms. Techniques like Query2doc (Q2D) and HyDE create contextually relevant paragraphs to append to the original query, helping to address issues with short, ambiguous, or imprecise queries.
However, as Zabir Al Nazi and colleagues from the University of California Riverside caution:
"LLMs can generate plausible-sounding but inaccurate or fabricated biomedical terms during query expansion. These incorrect expansions may shift the original intent of the query or introduce clinically invalid terminology".
To address this, grounding LLM prompts in trusted sources like UMLS, MeSH, or SNOMED CT can provide a safeguard. For instance, the BMQExpander framework, which integrates UMLS ontologies with LLMs, demonstrated a 6.5% performance improvement over the best non-grounded baselines.
Additionally:
"Incorporating precise definitions and reliable external knowledge into the prompt context enhances LLM understanding, improves reasoning quality, and reduces the likelihood of factual errors", note Zabir Al Nazi et al..
Best practices for LLM-based query expansion include using Chain-of-Thought prompting (e.g., adding a suffix like "Give the rationale before answering"), applying weighted expansion to keep queries anchored to their original intent, and pruning semantic graphs to focus on meaningful relationships like child, parent, and synonym connections.
How to Implement Query Expansion: Step-by-Step
Implementing query expansion in biomedical research involves a structured approach to ensure efficiency and precision. The process can be broken down into three key stages: preparing your queries, using ontology-based expansion, and merging sparse and dense retrieval models. You can also use tools like the Sourcely Knowledge Base to streamline your literature search. Let’s walk through each step.
Preprocessing Your Queries
The first step is to clean and standardize your query. This involves breaking it into individual words (tokenization) and removing stop words like "what", "or", "are", and "by", which don't add biomedical meaning. To improve retrieval accuracy, apply the Porter stemmer, which reduces words to their root forms. Research shows this method slightly edges out alternatives like the Krovetz stemmer in improving Mean Average Precision (MAP).
If you're using PubMed, note that its system often expands queries automatically. For tools like MetaMap, it’s important to remove stop words and punctuation beforehand to generate better candidate terms. Since preprocessing tends to prioritize recall over precision, consider reranking your results afterward using TF-IDF scoring to maintain the quality of your top results.
Applying Ontology-Guided Expansion
After preprocessing, map your cleaned query terms to medical vocabularies like MeSH or UMLS. Tools like MetaMap can identify clinical concepts in your text and link them to the UMLS Metathesaurus for query expansion. When working with MEDLINE-indexed documents, enable the "explosion" feature - this ensures your search captures documents tagged with more specific terms related to your main concept.
For better performance, combine ontology-based expansion with Pseudo-Relevance Feedback. Use a linear combination formula to balance the scores from both methods:
$L.C. = \alpha \times Score1 + (1 - \alpha) \times Score2$,
where $\alpha$ is adjusted between 0.1 and 0.9. This fine-tuning helps you strike the right balance between the two approaches. Once your query has been expanded using ontologies, integrate it with retrieval models for optimal results.
Combining Sparse and Dense Retrieval Models
The final step involves integrating your expanded query terms into both sparse and dense retrieval models. Use sparse models like BM25 alongside dense models such as BioBERT to refine your search. Studies show that combining query expansion techniques through linear combination can boost retrieval performance by 21.06% compared to baseline models.
When applying Pseudo-Relevance Feedback with Lavrenko's relevance model, adjust parameters like fbDocs, fbOrigWeight, and fbTerms to enhance accuracy. Consistently use the Porter stemmer during indexing and preprocessing to ensure term compatibility. This hybrid method, combining sparse and dense models, has consistently outperformed standalone approaches, with linear combinations achieving a 7.12% improvement over earlier biomedical retrieval studies.
Tools for Biomedical Query Expansion
AI-powered platforms and programmatic methods play a key role in improving the effectiveness of biomedical literature searches. These tools address challenges like vocabulary mismatches and broaden the semantic scope of queries, as discussed earlier.
Using Sourcely for Query Expansion
Sourcely leverages AI to take biomedical query expansion beyond basic keyword searches. By pasting paragraphs (up to 300 characters on the free plan), the platform identifies related biomedical concepts. It draws from a massive database of over 200 million scholarly papers, including sources like PubMed, PMC, Semantic Scholar, arXiv, CrossRef, and CORE.
One standout feature is the Deep Search functionality, which is especially helpful for handling complex biomedical queries. When standard search engines return generic results, this tool pinpoints specific peer-reviewed literature. You can refine your query expansions further by using advanced filters. These filters allow you to include or exclude particular keywords, focus on specific proteins, genes, or drugs, and even narrow results by publication year or peer-reviewed status.
"One of the limitations of databases like Google Scholar is that they let you search using only keywords. But what if you want to search using whole paragraphs or your notes? Sourcely is an AI-powered app that will let you do that." - Mushtaq Bilal, PhD, Postdoctoral researcher
Another useful feature is "Chat with Sources", which provides credible summaries and lets you quickly verify the relevance of your search results. This feature helps assess if the results align with specific research goals. Additionally, Sourcely supports citation exports in multiple formats, including APA, MLA, Chicago, Harvard, and IEEE. Pricing starts at $19 per month (billed annually) for up to 30 deep searches, with higher-tier plans available for more extensive needs.
For those who prefer a more technical, hands-on approach, the next section covers integrating PubMed APIs and UMLS.
Integrating PubMed APIs and UMLS
Programmatic methods offer a high degree of precision for query expansion, complementing AI-driven tools. With PubMed's E-utilities and the UMLS REST API, users can implement a manual query expansion process.
The process begins with UMLS's /search endpoint to obtain standardized Concept Unique Identifiers (CUIs). Once you have a CUI, you can use endpoints like /content/{version}/CUI/{CUI}/relations or /descendants to retrieve related terms, synonyms, and narrower concepts.
Next, you can send the expanded list of terms to PubMed's ESearch to retrieve PMIDs. Use ELink with cmd=neighbor_score to find similar articles, and rely on ESpell to correct any typos. Including an API key increases the rate limit from 3 to 10 requests per second. Additionally, setting usehistory=y in ESearch allows you to store results on NCBI's server for multi-step expansions using the WebEnv and query_key parameters.
Studies indicate that combining UMLS Metathesaurus knowledge with large language model (LLM) capabilities can boost retrieval effectiveness (measured by NDCG@10) by as much as 22.1% compared to standard sparse baselines.
Measuring Query Expansion Performance
To gauge whether query expansion techniques are effective, it's essential to evaluate them using standardized metrics. In the biomedical research field, specific benchmarks and metrics are widely used to objectively measure retrieval performance.
Performance Metrics for Query Expansion
One of the most commonly used metrics is Mean Average Precision (MAP). MAP assesses overall retrieval performance by calculating the average precision scores across multiple queries. This provides insight into how effectively a system retrieves relevant documents, whether for passages, aspects, or full documents. A higher MAP score indicates better performance, especially in ranking relevant results early in the list.
Another key metric is the F-measure, which balances precision and recall using the harmonic mean formula:
[ F = \frac{2 \times (precision \times recall)}{precision + recall} ]
This metric is particularly helpful in Boolean search systems like PubMed, where understanding the trade-off between retrieving more documents (recall) and maintaining accuracy (precision) is crucial.
For evaluating query expansion and keyword boosting, Inferred Normalized Discounted Cumulative Gain (infNDCG) is the go-to metric used in TREC Precision Medicine tracks.
Additionally, Mean Rank Precision (MRP) evaluates precision at specific cut-offs, such as P@5, P@10, or P@20, focusing on the performance of top-ranked results. While query expansion often improves recall by increasing the number of retrieved documents (e.g., from 1,174 to 3,163), it can sometimes reduce precision among top-ranked results.
"Query expansion is generally considered as a recall-favoring technique (as opposed to precision-favoring) that often results in a large number of additional retrievals." - Springer Nature
These metrics provide a robust framework for assessing and comparing different query expansion methods.
Benchmark Comparisons
Standardized benchmark datasets are critical for testing and comparing query expansion techniques. The TREC Genomics Tracks from 2006 and 2007 are considered a gold standard. They include biological queries (topics) and corresponding MEDLINE citations, totaling around 160,000–162,000 documents. Another widely used dataset is the OHSUMED test collection, which contains 348,566 MEDLINE references.
In 2011, researchers Sooyoung Yoo and Jinwook Choi from Seoul National University College of Medicine evaluated query expansion on the OHSUMED dataset. Using Local Context Analysis with a rank normalization reweighting method, they applied 15 terms from 50 pseudo-relevant documents. This approach led to a 12% improvement in MAP over the unexpanded baseline score of 0.2163. The results were confirmed as statistically significant (p < 0.05) using a paired t-test. Their findings highlighted that co-occurrence features outperformed frequency-based ranking in clinical query systems.
Different query expansion techniques yield varying results depending on the dataset. For example, while MetaMap (UMLS) often underperforms when used alone (achieving a MAP of 0.1685), combining it with other methods generally improves outcomes. Linear combinations of independent expansion techniques tend to deliver better results overall.
| Expansion Technique | Benchmark Dataset | Primary Metric | Performance Impact |
|---|---|---|---|
| Automatic Term Mapping (MeSH) | TREC Genomics 2006/2007 | F-measure | Showed improved performance; largest improvement noted |
| Lavrenko's Relevance Model (PRF) | TREC Genomics 2007 | MAP | Better results when combined with other methods |
| MetaMap (UMLS) | TREC Genomics 2007 | MAP | Underperforms alone; improves in combination |
| PubMed Dictionary + PRF | TREC Genomics 2007 | MAP | 21.06% improvement over baseline |
| LCA + Rank Normalization | OHSUMED | MAP | Achieved 0.2422 (12% improvement over baseline) |
These benchmarks and metrics highlight the varied performance of query expansion techniques, showcasing their strengths and limitations in improving retrieval accuracy.
Conclusion
Query expansion plays a crucial role in biomedical research by addressing vocabulary mismatches between user queries and the vast array of published literature. By incorporating strategies like synonym addition and term reweighting, methods such as pseudo-relevance feedback and dictionary expansion have significantly improved search outcomes. For instance, benchmark tests have shown an increase in the F-measure from 0.047 to 0.398 through these techniques. Combining multiple expansion methods has proven even more effective, with researchers reporting a 21.06% improvement over baseline search results.
Modern tools are making this process easier and more efficient. Platforms like Sourcely automate query expansion by identifying relevant sources and suggesting citation placements, helping researchers navigate complex biomedical terminology with less manual effort.
"Query expansions expand the search query, for example, by finding synonyms and reweighting original terms. They provide significantly more focused, particularized search results than do basic search queries." - BMC Bioinformatics
This guide has explored ontology-based, neural, and LLM-driven methods, each designed to tackle specific challenges in biomedical information retrieval. The future of literature search in this field lies in hybrid approaches, blending traditional methods like ontologies (MeSH, UMLS) with the capabilities of large language models (LLMs). These combinations promise both high precision and broad recall, making them invaluable for tasks like systematic reviews and background research. Whether you choose manual techniques or rely on specialized tools, adopting query expansion methods can significantly improve the accuracy and relevance of your search results.
FAQs
When should I use ontology-based vs neural vs LLM query expansion?
In biomedical research, selecting the right approach depends on your goals and the nature of your search. If precision is key, ontology-based methods are ideal. These rely on structured resources like medical ontologies to deliver highly accurate, knowledge-driven results. For queries that involve more complexity or ambiguity, neural models excel by capturing contextual and semantic subtleties. Meanwhile, LLMs (Large Language Models) are perfect for flexible, context-aware expansions and dynamic interactions, making them a great fit for exploring diverse datasets. Each method serves a distinct purpose, from delivering pinpoint accuracy to handling intricate, adaptive searches.
How can I prevent LLM expansions from adding wrong medical terms?
To ensure accurate medical terminology in LLM expansions, rely on domain-specific knowledge sources like medical ontologies (e.g., UMLS). These resources provide a structured framework to guide the process and reduce errors. It's also helpful to clarify or refine user inputs to avoid ambiguity, ensuring the system interprets queries correctly. Additionally, validating terms against trusted ontologies can further enhance reliability. By narrowing the scope of generated terms, you can maintain precision, especially in biomedical search and retrieval tasks.
What’s the simplest way to combine BM25 and BioBERT for better search?
The most straightforward way to combine their scoring systems is by using a weighted aggregation approach. BM25 focuses on assessing relevance based on lexical matches, while BioBERT creates semantic embeddings to capture contextual meaning. By combining these scores through a weighted function - where a parameter determines how much weight each model carries - you can strike a balance between semantic understanding and lexical relevance. This method improves the quality of biomedical search results.